Back to Forensic Science
Contents
- Development
- Background
- STRs
- DNA Profile
- DNA Match
- DNA Match Probability in CODIS-13
- Short Tandem Repeat DNA Database
At fixed spots on a chromosome’s DNA are genes, providing the blueprint for synthesizing proteins. But at other fixed spots are strange repeating sequences of DNA letters that differ from person to person. DNA Profiling is based on these repeating sequences.
Development
Sir Alec Jeffreys developed DNA profiling and used it to solve two rape-murders in a small country town in England in 1985 and 1986. A suspect had confessed to one of the crimes. But DNA profiling established his innocence and then was used to find the perpetrator.

Background
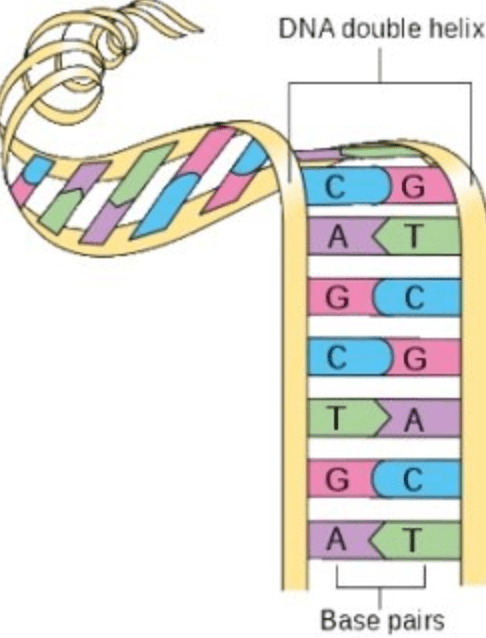
- A chromosome is a long string of DNA, which can be represented by a string of the letters A, C, G and T
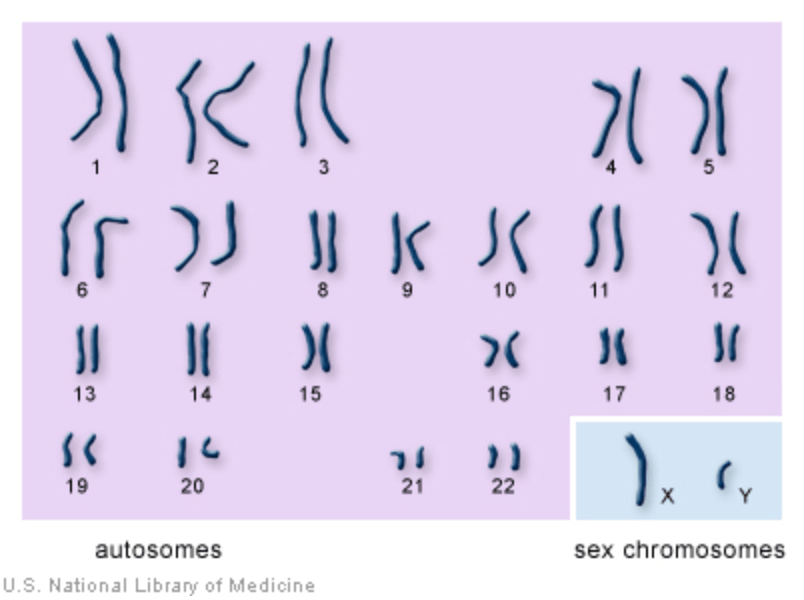
- Human beings have 23 pairs of chromosomes
- A gene is a sequence of DNA that
- occupies a fixed position (locus) on a chromosome
- directs the synthesis of a protein.
- For example, the HBB gene codes for the protein Β globin and is located on chromosome 11, extending from locus 5,225,464 to locus 5,229,395. It is represented by a sequence of 3,932 occurrences of the letters A, C, T, and G.
STRs
- Not every DNA sequence on a chromosome is a gene. Among sequences that are not genes are STRs: short tandem repeats.
- For example, at position D7S280 on chromosome 7 the letters GATA are repeated 6 to 15 depending on the person. The DNA sequence of alternating red and green below has 12 repeats of GATA:
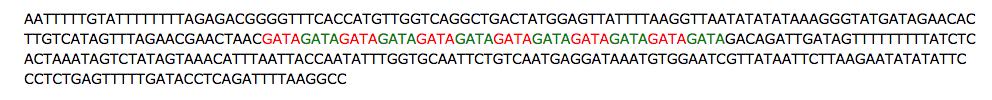
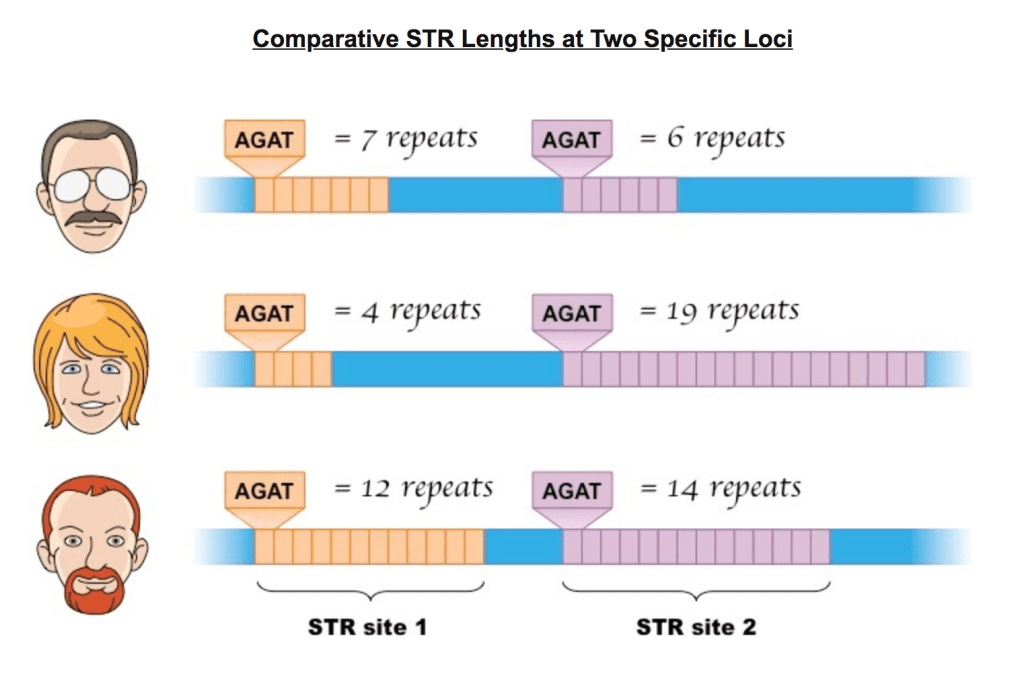
- The number of repeats at a given STR location varies from person to person
- In the diagram are the repeats for two STR sites.
DNA Profile
- A DNA Profile consists of a list of certain STR locations coupled with the number of repeats at each location.
- Below are two CODIS-13 DNA Profiles, one for Forensic Unknown, the other for Candidate Offender.
- The two profiles are a partial match, indicating a genetic relationship.
- The numbers:
- A single number is for both chromosomes
- Two numbers are for each chromosome
- A decimal is a partial repeat
DNA Match
- Before 2017 the FBI’s Combined DNA Index System (CODIS) used 13 STR locations. CODIS now uses 20.
- Two CODIS-20 DNA profiles match if they have the same number of repeats at all 20 STR locations.
- If two DNA profiles don’t match, the profiles are of different people
- If two DNA profiles match then, except for identical twins, it’s astronomically likely that the profiles belong to the same person
- In general, a laboratory error, contamination, or fraud is more likely than a false positive.
DNA Match Probability in CODIS-13 (excluding identical twins)
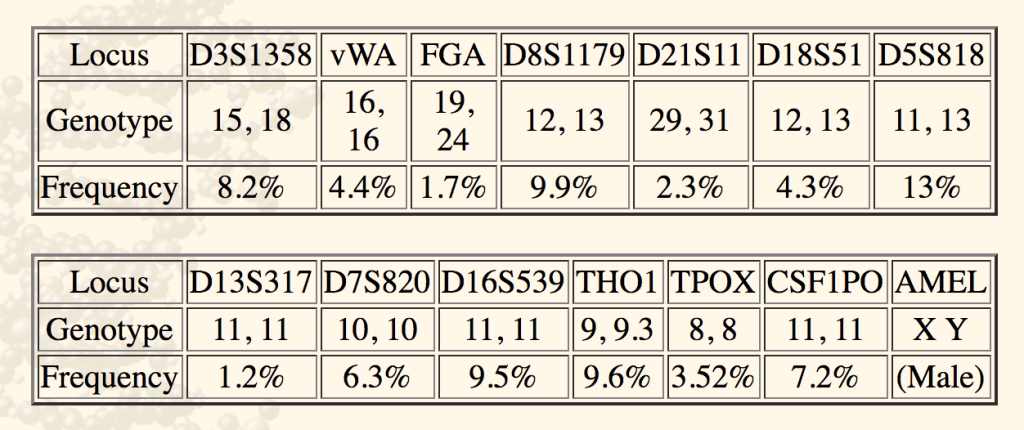
- Per the Special Conjunction Rule, the probability of another person’s profile randomly matching this one is 0.082 * 0.044 * 0.017 * 0.099 * 0.023 * 0.043 * 0.13 * 0.012 * 0.063 * 0.095 * 0.096 * 0.0352 * 0.072 = 0.00000000000000001364.
- The Special Conjunction Rule assumes STR’s are independent, e.g that TH01 = 9,9.3 doesn’t affect the probability that FGA = 19,24.
- STR Analysis (Wikipedia)
- The true power of STR analysis is in its statistical power of discrimination. Because the 13 loci that are currently used for discrimination in CODIS are independently assorted (having a certain number of repeats at one locus does not change the likelihood of having any number of repeats at any other locus), the product rule for probabilities can be applied. This has resulted in the ability to generate match probabilities of 1 in a quintillion (1×1018) or more. However, DNA database searches showed much more frequent than expected false DNA profile matches. Moreover, since there are about 12 million monozygotic twins on Earth, the theoretical probability is not accurate.
- View Special Conjunction Rule
- Assuming STRs are independent and setting aside identical twins, the probability of another DNA profile matching the one above by chance is about one in 73 quadrillion
- 73 quadrillion is 8 billion (the world population) times 9 million.